RELION
A powerful biomacromolecular structure determination package for CryoEM applications.
RELION requires an environment module
In order to use RELION, you must first load the appropriate environment module:
module load relionThis application is available in Open OnDemand.
RELION is a powerful hybrid parallel software package with GPU support for CryoEM structure determination. The program uses an empirical bayesian approach to determine biomacromolecular structures from 2D and 3D datasets.
Using RELION on HPC Systems#
RELION can be used both interactively via the GUI and non-interactively via a SLURM Script. Non-interactive mode is recommended for long-running jobs.
RELION can be accessed as an interactive app or through the command-line.
Accessing RELION as an Interactive App#
First, you must upload your data onto the HPC. Or, if your files are small, you can upload them directly using Open OnDemand
- Log into Open OnDemand
- In the top navigation bar, select "Interactive Apps" and then from the drop down select, "RELION".
- Select a Slurm Account/partition that includes GPU nodes (either
backfill2,gpu_q, or an owner account/partition.). - Complete the remaining sections of the job submission form.
- Ensure that the "Amount of Memory" is at least 128 GB and that you select a minimum of one GPU.
- Under "Path to RELION Project Directory", type the full path to the directory you plan to use for your RELION project. RELION will be launched from within this directory. If you do not specify this, the default will be your home directory.
- Click "Launch" to queue your job.
- When your interactive job has started, click "Launch RELION".
- A new tab will open, and the RELION GUI will launch shortly after.
Motion Correction with UCSF MotionCor2#
When performing motion correction using UCSF MotionCor2 on the HPC system, the following settings must be configured correctly:
From the top left menu.
* From the I/O Tab Configuration in the top left corner:
- "Write Output in float16": Set to "No".
- "Save Sum of Power Spectra": Set to "No".
* From the Motion Tab Configuration in the top left corner:
- "Use RELION's Own Implementation": Set to "No".
- "Use UCSF MotionCor2 Executable": With the previous option set to "No", you can specify the executable path. Enter the following path:
/gpfs/research/software/motioncor2/MotionCor2_1.6.4_Cuda118_Mar312023.
These settings ensure that motion correction is handled effectively and without conflicts.
Accessing RELION through the Command-Line#
To use RELION, you only need to load the RELION module by the command module load relion.
This will load the current default version of RELION into your path.
From here, you can run any of the RELION programs, or you can run the relion command itself to start the GUI.
The RELION module automatically loads the GNU 8.5, OpenMPI 4.1.0 and CUDA 11.8 modules as these are required
to run the program in parallel with GPU support.
Starting the RELION GUI#
Note
You will need to either use Open OnDemand or start a graphical SSH connection to use the Relion GUI
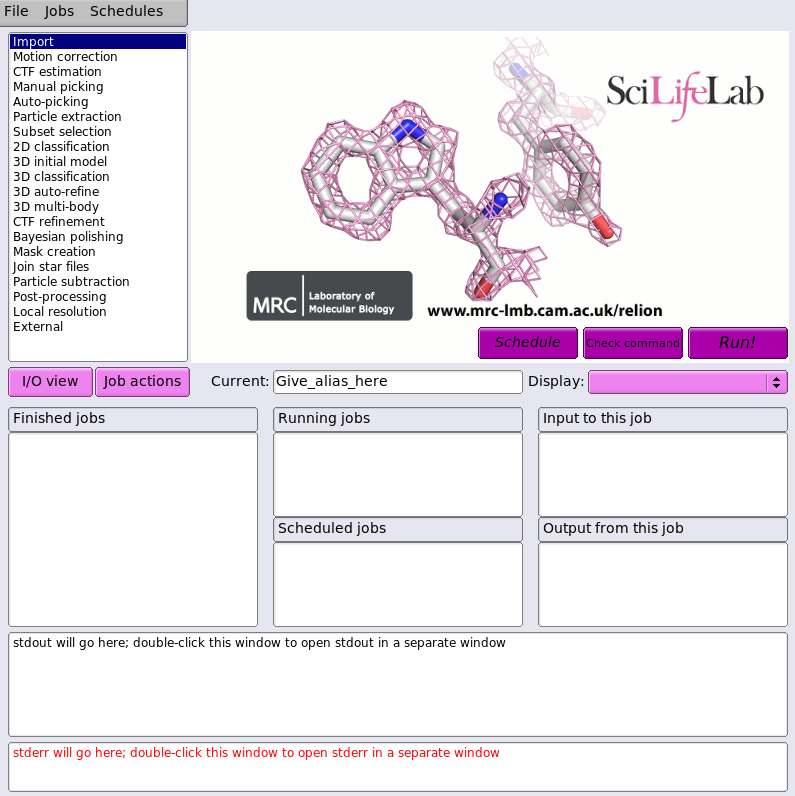
To start the RELION GUI, from which you can submit jobs, first navigate on the login node to a RELION work directory you have created. Then run the RELION command:
This will start the RELION GUI as shown below:
Example Submit Script for RELION#
The following submit script is an example which uses the relion_refine_mpi subprogram with data from the
shiny_2sets.star tutorial dataset. This script may require significant modification for optimal performance
on the HPC system and may require modification for use with other RELION subprograms other than the
relion_refine_mpi program shown in the example. However, this is a good starting point. This test script was
prepared, in part, with the assistance of UF Research Computing.
Once you have prepared your job script, be sure to save it as a filename, for example "myrelionjob.sh."
You can then submit it via sbatch myrelionjob.sh.